Our research is to understand the epigenetic regulation of transposable elements and how their dysregulation contribute to the generation and development of blood cancers. In particular, we investigate their roles as gene regulators and triggers of anti-tumour immunity in blood cancers. Our long-term vision is to address how we can utilise transposable elements as a gateway to develop new and effective treatments for cancer.
Background
Transposable elements (TEs) are mobile DNA segments that have expanded within the human genome throughout evolution. TEs have evolved cis-regulatory sequences to exploit the host cellular machinery to promote their own transcription and replication. Whilst the vast majority of TEs in the human genome have been rendered immobile due to mutation, many have retained their regulatory function. Given their selfish nature, TEs have dispersed vast amounts of cis-regulatory sequences and transcriptional units across the genome, providing an abundant source of transcriptional modulatory elements.
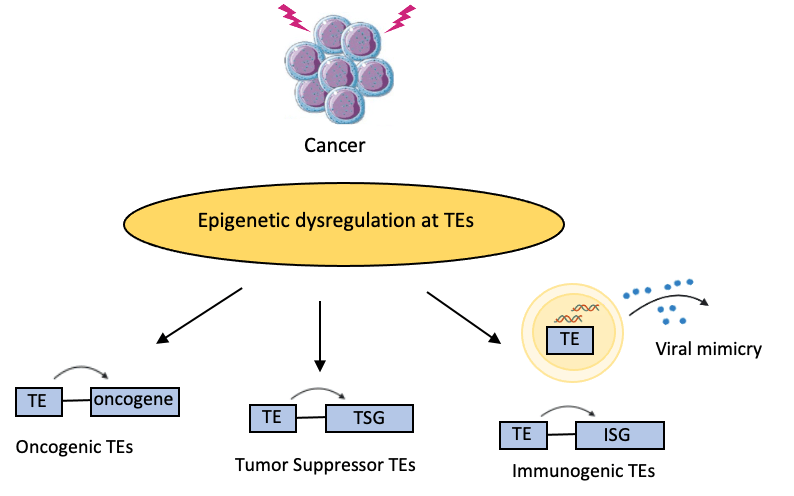
Epigenetic dysregulations are hallmarks of cancer and that provides a particularly fertile ground for TE activation. In fact, we revealed the first examples of TEs activated as oncogenic enhancers in acute myeloid leukaemia, which conferred a proliferative advantage to the cells (Deniz et al, Nat. Comms., 2020). In contrast, genome-wide epigenetic changes in cancer cells could also potentially activate TEs that regulate the expression of tumor suppressor genes, leading to loss of cancer cell fitness; or trigger anti-tumor immune responses leading to the destruction of cancer cells via “viral mimicry”.
Our Goal
Using AML as a model system, the overarching goal of the Deniz Lab is to comprehensively delineate the molecular and cellular functions of TEs in cancer genome and characterise the epigenetic mechanisms that define their diverse activities. To achieve this aim, we will:
- Identify TE-derived cis-regulatory DNA elements and determine their transcriptional effects in AML
- Assess how these regulatory TEs modulate cancer cellular phenotypes
- Dissect the role of TE-mediated immune response in AML.
- Identify driver epigenetic events in TE regulation
Our Methods
In the Deniz lab, we employ high-throughput techniques to gain a genome-wide view of transposon activation/regulation in cancer. This includes ATAC-seq (bulk and single-cell levels), RNA-seq (long/short-reads, bulk and single-cell levels), ChIP-seq, CUT&RUN, CUT&Tag, Micro-C, targeted bisulfite sequencing and EM-seq. The bioinformatic analysis of the genomics data allow us to test our hypotheses and provide specific candidates to perform functional assays using CRISPR/Cas9-based genome and epigenome editing. We perform these experiments in cell lines and primary cancer samples and use xenograft models to decipher the impact of transposons in cancer genomes.
We are also excited to learn about and employ Mass spectrometry-based approaches to gain insights into the regulatory mechanisms of transposons and explore their contribution to adaptive immunity in cancer.

